cm3020 Lecture Summaries: Topic 02
Notes from the lectures in cm3020 Topic 02: Automated Scientific Discovery
Week Six lectures
6.101 Discovery Science
Guest lecture from Prof. Ross King (Cambridge), lead on the Robot Scientist project.
The application of AI to science is called scientific discovery. Reviews the cyclic nature of AI research (hype booms and busts).
AI systems have superhuman scientific reasoning powers. They can:
Flawlessly remember vast numbers of facts
Execute flawless logical reasoning
Execute near optimal probabilistic reasoning
Learn more rationally than humans
Learn from vast amounts of data
Extract information from millions of scientific papers.
Argues that AI is a great application area for science:
Scientific problems are abstract, but involve the real world.
Scientific problems are restricted in scope: to be good at science AI systems don’t need to know about anything else but science.
Nature is honest, no malicious agents (unlike economics, warfare).
Finishes with a plea to stop developing ads and save the world!
6.103: The Robot Scientist
Discusses concept, motivation, and architecture of a robot scientist. Introduced by Ross King in 2004. Demonstrated that it was possible to automate scientific disovery, with the system ‘Adam’.
A robot scientist is an AI system capable of originating its own experiments, physically executing them, interpreting the results, and then repeating the cycle.
Shows the architecture:
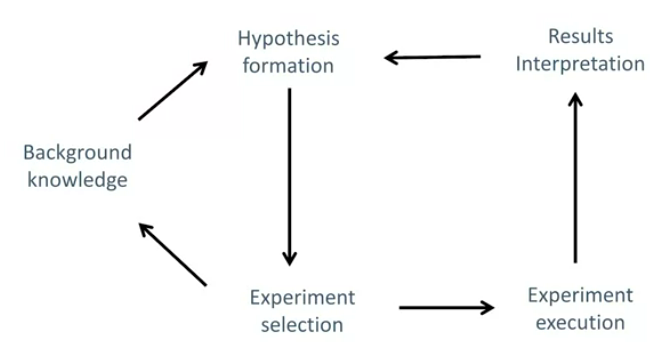
RS can infer from its knowledge base a new hypothesis - something not already encoded in its knowledge base. This hypothesis is a possible explanation of the available phenomenon, not necessarily true. It then tests this hypothesis, it can select and run experiments in parallel, and then interpret the results. A new hypothesis is formulated, and so on.
This is an implementatin of the hypothetico-deductive method of scentific disovery:
A procedure for the construction of a scentific theory that will account for results obtained through direct observation and experimentation and that will, through inference, predict further events that can then be verified or disproved by empirical evidence derived from other experiments.
Several sciences involve experimentation. Not all can be automated. This is still in its infancy, but has immense potential to speed up scientific discovery.
Why robot scientists? 1) If we can create a machine that does science, then we understand what science is.; 2) Production of data outpaces knowledge production, we don’t have resources to crunch the data or papers produced, robot does…; 3) Robots can record and share all aspects of discovery, helping open science; 4) not enough human scientists.
Humans are great at developing a deep understanding of phenomena, making conclusions based on small data, and leaps of imagination. Computers better at processing big data and making unbiased, accurate logical inferences. Application of RS is limited to certain scientific areas amenable to automation.
6.201: History of RS
Many believe it’s not possible to automate scientific discovery, but there is a history of autonomous discovery.
Argument: eg Popper “every discovery contains an ‘irrational element’, or a ‘creative intuition’” (1961, Logic of Scientific Discovery).
Dendral was one of the pioneering systems that proved Popper wrong. Developed in Stanford in the 1960s, aimed to study hypothesis formation and discovery in science. Task: help organic chemists in identifying unknown organic modules, by analysing mass spectra and using knowledge of chemistry. Several systems derived from dendral.
Bacon was another system, developed in the 80s. Searched through a problem space of algebraic terms, using operators to generate new terms from old ones. Led to a rediscovery of Kepler’s third law, and a variety of numeric relations.
Mentions several other papers giving examples of automated discoveries. Then turns to the Robot Scientist project, introduces phases of the project. Main phase focused on yeast functional genomics. Most functions of the genes remain unknown. Tried to determine gene functions by growth experiments.
Had a background knowledge model, the yeast metabolic model. Could access bioinformatics databases. Hypothesis generation was done through abductive reasoning, and bioinformatic techniques. Experiments were designed through Random Forest and other ML tools.
You can see Adam in this video
Adam was the first machine to autonomously discover novel scientific knowledge: hypothesise, and experimentally confirm it.
The next RS was named Eve, worked on drug design for tropical diseases. Parasites are neglected by pharma, so RS are good here. Knowledge was encoded as graphs.
Next stage was AdaLab, European project, application domain was systems biology. Models are highly complex, so good area for RS. Produced many ML tools and models. Results were important to cancer and aging results.
Current stage is Genesis. Hardward can execute 10,000 closed-loop cycles in parallel.
Other examples include the Automatic Statistician aiming to automate data science, see https://doi.org/10.1007/978-3-030-05318-5%5F9.
To date, discoveries are on a modest scale, but the aim is to build systems that are capable of significant discoveries.
6.301: Ethical and Legal Issues
Focusing on ethical and legal issues related to the case study, and laboratory automation. Roboethics covers ethical issues related to robot design, operation and use. Can we design robots to act ethically? How should people treat robots and vice versa. Who is responsible if the robot causes harm. Are there risks to create emotional bonds with robots?
RS project criticised for replacing scientists. General issue of robot work replacing human. Lab automation is replacing technicians? Claims that it is not an issue in this area. The goal: To automate repetitive tasks, or tasks that must be performed in hazardous environments. Example: during pandemic labs were shut, can we develop remote autonomous laboratories?
Looks at collaboration: AI systems better at processing large volumes of data, reasoning, and parallel tasking. AI systems work 24/7. Humans better in: high-level planning, making decisions in uncertain, non-deterministic and partially observable environments. Target: human-robot collaboration.
Another issue: Bias and mistakes. EG medical datasets are biased, not enough data from minority groups. So medical systems can make biased decisions as a result. Bias has to be made explicit and mitigations implemented. AI systems can make mistakes. In RS, all experiments are replicated many times.
Legal issues: Existing laws not developed with AI in mind. EG Adam could not sign the publication contract with Nature, so couldn’t be a co-author.
Safety: AI system must be designed in a way that do not pose a threat to humans. Adam was placed in an enclosure. Eve had sensors, if a person approached dangerous equipment the robot would shut down automatically.
Finishes by saying that lab automation less prone to ethical implications than other areas of AI, but the issues are critical for the wider field.
Week Seven Videos
7.101 Introduction to Rational Agents
This video summarizes Russell Norvig Chapter 02: Intelligent Agents.
7.104 Multi-Agent Environments
Weirdly focuses on games and adversarial search, even though that’s not the context of the robot scientist. Introduces the prisoners’ dilemma, and robot football, then mentions Alpha Go. Not clear what the point of this lecture is.
7.107 Overview of the Toy Robot Scientist System
Introduces the toy robot scientist. It is designed to assist in drug discovery. It is a multi-agent system. It has a knowledge agent that can access a data set and update its knowledge model. A hypothesis generation agent that can create hypotheses from the knowledge base. A planning agent can design experiments based on the hypothesis. It passes a protocol to the robotic lab agent which can run the experiment. the lab passes the data to the results interpretation agent. This agent can decide if the hypothesis is confirmed. New data and knowledge about confirmed or rejected hypotheses are added to the knowledge base.
This could be enhanced by:
A meta-analysis agent, collecting information about runs, results and compares performance of other agents.
A collaboration agent, that supports a team of scientists, requiring orchestration of the goals, knowledge basis, complex scheduling of experiments and sharing outputs.
A monitoring agent, monitoring and recording everything that’s happening. How long did everything take? Are there enough resources?
An external services agent, an interface for external users. EG users could submit their hypothesis for testing and conclusions by the results interpretation agent.
A model agent can work not only with data and rules, but also executable computational models, not clear what this is really.
An explanation agent can explain to users why and how these conclusions were made and not others. Work on this is still nascent.
7.201 Crash Course in Knowledge Representation
Introduces how knowledge can be represented and encoded in a machine processable way.
There are dedicated knowledge-representation languages, that allow inferences. There are different logics: description logic, propositional logic, first order (predicate) logic, higher order logics, temporal, modal etc. Different logics feature a different balance expressive power and reasoning complexity. Lecture looks at first order logic, and description logic, and RDF and prolog.
DL is decidable, so an inference engine can arrive at a conclusion, not true of predicate logic.
Presents an example of a passage of text about the degree programme. Two step process to extracting knowledge from it. First we extract entities, then we model relations.
Examples of relations in the example: enrolled(student, module); raises(student, query); is_a(BSC CS, undergrad programme).
Shows a knowledge graph with the relations. Introduces RDF:
provides a model for describing resources (objects, entities)
defines a resource as any object that is uniquely identifiable by a URI
Resources have properties that link resources to other resources (ie relations, or object properties)
Resources can be linked to some values (like strings, numbers), data properties
RDF encodes knowledge in triples. The logic is a description logic, making it easy for reasoning agents to work with.
There are different schema available for RDF, introduces RDFS and Turtle (.ttl), the easiest to read for humans.
rdfs:Class is used to classify resources eg:
ex:Module rdf:type rdfs:Class,ex:ML rdf:type rdfs:Module.ais a shorthand forrdf:typeso these can be written asex:ML a rdfs:Moduleex:being a placeholder for the namespace.rdfs:Propertyis used to add relations to classes:ex:raises a rdfs:Property. Statements can be generated from a knowledge model.Knowledge models can be extended or integrated with other knowledge models.
Definitions in rdf are global and unique. Resources can be explicitly linked or imported into a knowledge model.
Integration can be achieved by linking entities defined in different knowledge models.
Gives example application of knowledge model, an agent for classifying queries as academic or non-academic and deciding where to escalate them.
7.203 The Knowledge Base
Introduces the domain of drug discovery. Takes ten years to bring new drug to market. Starts with a library screening. A library is a collection of chemical compounds. Screening is running experiments with chemical compounds in a lab. Hits are compounds that show biological activity. Next we have to confirm the hit, if we do we have a lead.
Toy RS is focused on identifying a lead. If we could speed this up through automation process will be much better!
How can we represent the knowledge? Molecular structure consists of atoms and bonds that connect them. Lots of encoding schemes for these. Common to represent these as a table for ML purposes. Eg:
| Compound | Molecular Weight | Hydrophobicity (LogP) | Hydrogen Donor | Hydrogen Acceptor | Heavy Atom | Aromatic Ring |
|---|---|---|---|---|---|---|
| Aspirin | 180.16 | 1.31 | 1 | 3 | 13 | 1 |
| Cetirizine | 388.9 | 3.15 | 1 | 27 | 2 |
First we design a conceptual model of our domain, blue we know from the table, yellow is additional facts:
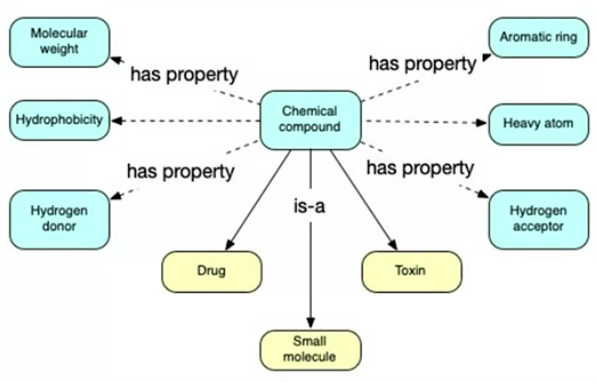
Small molecules are more likely to have an effect as they can penetrate cell membranes. Common domain knowledge is not needed to rediscover through ML. How do we add new information to models to improve the learning in traditional ML? Can only do it if it fits the data format. Complex, multi-layered domain knowledge is hard to do in tabular or database form and hard for ML to replicate. This is where logic encoding comes in.
How do we represent the toy domain knowledge? Encode directly in Python? Tricky. Encode in RDF? OK. Use Protege to encode in RDF and export to python? Better, reasoners can check logical consistency and infer new facts. Encode in predicate logic in Prolog, good but requires prolog knoweledge.
7.204: Constructing the Knowledge Model
Looks inside a knowledge agent. Uses Protege. Useful tool that forces good clean encoding. Exports to rdf and turtle.
Walks through starting a new ontology from scratch in Protege, adding classes, instances, and data properties in the UI.
Shows reasoning with Fact++, and logical consistency checks.
Shows adding a Small molecule class which is a ‘defined class’, where there are constraints, here where molecular weight is below a threshold it will be defined as a small molecule.
Shows exporting to ontology syntax (eg turtle), just by saving the file.
7.205: Demo of Toy Robot Knowledge Model
Walks through main.py introduces the consolemenu library for easy terminal UI: https://github.com/aegirhall/console-menu
Walks through knowledge_model.py. Uses rdflib library (see github repo).
7.206: Constructing a Real Knowledge Model
Demonstrates some real knowledge models, vaguely. Demonstrates the CHEBI ontology.
Wanders through drugbank and ChEMBL (human created databases). Then the AdaLab ontology.
Week Eight Videos
8.101 Crash Course on Logic
Agents’ intelligence requires the ability to reason over the knowledge base. Knowledge has to be encoded in logic to enable this reasoning.
We’ll look at first order logic and prolog. Representing and encoding a statement in logic enables a logical inference engine to determine if the statement is true or false.
Introduces the basic logical operators, negation, conjunction, disjunction, implication and equivalence. Gives a very poor explanation in terms of a confusing venn diagram and truth table.
Introduces quantifiers, universal and existential.
Turns to Prolog, a language designed for solving problems involving objects and the relations between them.
We can express a relation like this: raises(pete, query_001) for “Pete raises a query with id query_001”. This is a clause or logical statement.
Instances like Pete start with a lower case letter, variables start with an upper case letter, predicates start with a lower case.
The inference engine can be used to ask questions of the knowledge base and draw inferences based on FoL. Mentions the similarity to relational databases, and the research area of inductive databases.
Shows an example of implication - students are classmates if they are enrolled in the same module. Shows the common issue that you also have to specify they aren’t the same person. Talks through prolog concepts:
The head established if the statements in the body are true. We use :- for implication, , for conjunction, ; for disjunction. Examples:
successful(X) :- rich(X) ; famous(X). Heresuccessful(X)is the head, rich or famous is the body.classmate(S1,S2) :- enrols(S1,M), enrols(S2,M), dif(S1,S2).
Questions we pose to a prolog engine are called goals. These can be complex statements.
Finishes with some contrast of first order logic and descriptive logic. FoL is to describe properties and relations among objects,Dl is to define and describe objects.
Inferences in DL are subsumption (if one class is a subclass of another) and classification (inferring class membership).
8.103 Types of Reasoning
Deduction: Conclusion of the consequent is guaranteed by the truth of the premises. Eg subsumption, inference about class-subclass structure. Classification, inference about what class an object belongs to.
Abduction: References Charles Sanders Peirce. Generates hypothesis.
Induction: Generalization of facts, not guaranteed. ML is based on induction.
Those will be what we focus on. Other types are mentioned…
8.105 Hypothesis Generation: Abduction
Deduction: demonstrates modus ponens, contrasts with abduction where the truth of the premises does not guarantee the truth of the conclusion. Abduction here seems to be the old (muddled) Peircean sense, not the modern use of the term (see the SEP entry.
Discusses the use of abduction in Adam. The bacground knowledge base is a metabolic pathways model of yeast - a graph representing chemical reactions. The graph is encoded in prolog.
Yeast has about 6,000 genes and their functions are similar to those in humans. It’s still unknown what genes are responsible for some of the reactions. It is possible that several genes are responsible for one reaction and one gene may be responsible for several reactions.
Adam considers the 6,000 genes and their combinations, what genes may control the biosynthesis of a chemical. Adam has access to bioinformatics databases. It can find what enzyme catalyzes the considered reaction, what genes are responsible for that catalyzer in other organisms, and homologous genes in yeast.
Adam abduces possible options, inserts candidates to its logical model, and runs a reasoner to check if the chemical can be synthesised.
8.107 Hypothesis Generation: Induction
Generalization of facts, not guaranteed to be correct. Associates ML with inductive reasoning, finds patterns in observations and generalizes them to new observations.
Gives some examples of induction in biological research.
8.201 The toy abduction reasoning agent
Main file is abduction_reasoning.py Uses the kanren library to perform the reasoning.
No real explanation of kanren (some is provided in the docs in the repo).
Week Nine Lectures
Cover planning, scheduling and analysing experiments.
9.101 Planning and Scheduling
Planning is a critical part of AI, at the core of applications such as manufacturing and robotics.
It is supported by dedicated languages, eg the Planning Domain Definition Language.
Classical planning deals with observable, deterministic situations. A planning task is a search problem: Given an initial state, a list of possible actions, and their results, find a sequence of actions that leads to the goal state.
We need to be able to represent the states and actions. Eg using a plane example:
Action: fly(p,from,to)
pre-condition: at(p,from) & plane(p) & airport(from) & airport(to)
effect: not at(p,from) & at(p,to)
Walks through a block moving puzzle, modeling it in prolog.
Planning in the real world is more complex, the environment may be partially observable, nondeterministic, or unknown. We may need to do contingent planning, monitoring and re-planning, and the world may not be closed. See Task Environments for more on this.
Scheduling adds constraints to sequential plans, time and resources. The typical approach is “plan first, schedule later”.
Mutli-agent planning is more complex, we may need to add an extra layer of planning and scheduling. Agents may share the same goal and the same representations, or not. They may need to co-operate or communicate.
9.103 Planning experiments
Not all formulated hypotheses can be tested. Some might depend on equipment or materials not present. Sometimes we need to test a related hypothesis. Eg we can’t experiment directly with genes, so proxies are used.
Gives an example of Adam outputting a set of hypotheses (encoded in prolog). The planning agent then needs to know the available equipment, materials and a knowledge base for designing experiments. An ontology was created for this, LABORS link provided.
Planning agent designs an experiment to see if eg behaviour changes in a yeast strain with a gene of interest removed.
Experimental knowledge bases are still largely added manually by scientists, as are protocols and parameters. EXACT is an ontology for experiment protocols. It needs to be translated to a lower-level machine processable protocol. No current standardised protocol across different manufacturers.
9.201 Toy Planning Agent
Walks through code in planning.py. Nothing particularly interesting.
9.301 Experimental Analysis
Starts by trashing human analysis as biased and prone to over-generalization.
Robot scientists produce a lot of experimental data. use ML to analyze the data.
Talks about the yeast example, RS used decision trees to decide if the result was significant.
Then talks about Eve, identifying drug leads for Malaria.
Week Ten Lectures
Week ten introduces current research projects of the lecturer.
10.101 Meta-analysis
A meta-analysis agent can collect information about previous results and compare them with performance of other agents to compare situations and what works best.
EG meta-machine learning is learning about ML algorithms and which algos perform well in which circumstances.
Meta-QSAR is such an agent for Eve. Lecture summarises some of the work in the published paper.
The work in the paper analysed which ML algorithm performed best over a range of datasets. It found that Random Forests performed best nearly half the time, but that means that there are over half of scenarios where it didn’t.
If we can prepare a meta-dataset - a dataset about our datasets, perhaps we can predict which ML algorithm will perform best based on features we engineer or identify about those datasets.
Results seemed promising, suggesting that meta-learning techniques can be employed to help select the best ML algorithm for a particular dataset.
10.103 Communication Agents
Looks at a communication agent developed for Eve. Introduces multi-agent environments - acting as a swarm (no communication), competitively, or cooperatively.
To cooperate agents need to share knowledge about the goal, world, and each other. For communciation we need content, a protocol for exchanging information, and a mode of communication.
Developed a protocol for Eve, SciCom, minimalistic, defines syntax and semantics for communicating requests for experiments, output of results etc.
Based on the Eve ontology, which defines a standard experiment. SciCom describes differences from the standard. Implemented in yaml
Support communication between people and robot scientists across different labs.
10.201 New projects
Introduces current research from the lecturer:
Project one - decision support system for the treatment of cancer. Develop a prototype AI system for the design of personalised cancer treatments. Focus is on chemotherapies (drug cocktails).
Challenge is that with single drugs cancers can evolve resistance quickly. Cocktails are better but there is a combinatorial explosion. AI is going to help select suitable candidates.
Project two - AMBITION - aims to provide researchers with continuous, remote access to AI/robotic augmented biomedical research capabilities. Pandemic showed the importance of biomedical research continuing inspite of social distance/lockdown measures.
Project is putting together a demo lab to show that remote, automated, closed-loop robotic experiments are possible. Not aiming to replace humans, rather to support real-time treatment adaptation supported by continuous testing of novel drug combinations on tumour material.