Jurafsky Manning Lecture Summaries: Topic 08 Information Extraction
8.1 Introduction to Information Extraction
The goal of information extraction systems is to find and understand limited relevant parts of texts. Typically we gather information from many pieces of text.
The goal is to produce a structured representation of relevant information:
relations (in the database sense)
a knowledge base.
We might be doing this to organize information so that it is useful to people; or to put information in a semantically precise form that allows further inferences to be made by computer algorithms.
IE systems extract clear, factual information (who did what to whom when?)
EG gathering earnings, profits, board members, HQ etc from company reports. Eg extracting from “The headquarters of BHP Billiton Limited, and the global headquarters of the combined BHP Billiton Group, are located in Melbourne, Australia”, the relation headquarters("BHP Biliton", "Melbourne, Australia")
Low-level IE is embedded now in applications like GMail, extracts eg dates. Often this kind of IE is simple, and based on regex and name lists.
A very important sub-task is Named Entity Recognition or NER, where we find and classify names in a text.
Finding a name involves finding its limits in the text. Once we’ve found those limits we can classify them (eg as Person, Organization etc).
The history of the task is reflected in the name, entities were supposed be discrete entities in the world, eg Stanford, but not sand, is an entity. Some entities have names, eg Chris Manning, but not all, eg the chair you’re sitting on. The idea is we’d pick out entities with names.
But that’s not quite what we’re doing now, which is more “easily distinguishable names or other things in text that we can pick out” (like dates, quantities, times).
The uses of NER:
They can be indexed, linked-off, etc.
Sentiment can be attributed to companies or products.
A lot of IE relations are associations between named entities.
For QA, answers often invovle named entities.
Often a sub-task in a broader task like QA, sentiment analysis…
8.2 Evaluating Named Entity Recognition
In a NER task we have an input sequence of word tokens, and we want to predict entities.
Eg in the sequence “Foreign Ministry spokesman Shen Guofong told Reuters” we want to predict that the two tokens “Foreign”, “Ministry” are one entity, “Shen” and “Guofong” are another, “Reuters” a third.
Standard evaluation is per entity not per token. So when we draw up a confusion matrix we do it on the entity basis.
So we have entity based precision and recall. But the details are tricky.
In standard contexts for precision and recall there is one grain size, ie one size of unit that you’re eg classifying (a document).
But the measure gets funny for IE NER when there are boundary errors, which are very common.
Take the entity “First Bank of Chicago”. Say our algorithm predicts “Bank of Chicago” as an entity, missing “First”. Now we have both a FP and a FN. It would have been better to select nothing.
Intuitively we should say that the NER recognizer was mostly correct, no? It identified an entity, got the right boundary correct, classified it correctly as an organization, it just missed one token. If the true label is ORG-(1,4) we got ORG-(2,4).
Some other metrics have been suggested, that give partial credit, eg the MUC scorer. But then you get complex questions of when to give partial credit. So most of the field haven’t gone there, and have stuck with simpler f1 metrics despite the issues.
8.3 Sequence Models for NER
The ML sequence model approach to NER looks like this:
Training
Collect a set of representative training documents
Label each token for its entity class or other (O)
Design feature extractors appropriate to the text and classes
Train a sequence classifier to predict the labels from the data
Testing, Classifying
Receive a set of testing documents
Run sequence model inference to label each token
Appropriately output the recognized entities
There are different encoding schemes that differ on how they represent entity boundaries.
The simpler is IO encoding (Inside/Outside), which just assigns a category label or ‘O’ if the token is part of the entity or not.
But how do you tell when one entity’s name ends and another entity begins?
We do that with IOB encoding where each class is prefixed with ‘B-’ for beginning of an entity, or ‘I-’ if it’s a continuation.
But that comes at a cost, instead of C+1 labels we have 2C+1 labels where C is the number of classes. This ends up slowing down encoding.
So some tools still use IO encoding (like the Stanford tool), since confusing situations don’t often occur. Often adjacent entities are different classes. IOB encoded systems are harder to train, and often get the hard problems wrong anyway.
Input IO Encoding IOB Encoding Fred PER B-PER showed O O Sue PER B-PER Mengiu PER B-PER Huang PER I-PER ‘s O O new O O painting O O Features for sequence labeling:
We use the current word (like a learned dictionary of words in each class).
Previous/next word (for context).
We might use POS tags, current, previous and next word POS tags.
So far we don’t have a sequence model, we can introduce label context (eg previous and perhaps next label), which then becomes a sequence model. EG it might be common for person entities to be more than one token, so we use the prior token’s classification as a person to predict that the next token is more likely to be so too.
Character subsequences of a word can be very useful for classification of entities. For example the substring ‘oxa’ might be a really strong indicator of the word being a drug name. A ‘:’ token might be a really strong indicator of a title. A word ending in ‘field’ could be anything, but is more likely to be a place.
Word shapes can also be helpful. We map words to equivalence classes that are simplified representations that encodes attributes such as length, capitalization, numerals, Greek letters, internal punctuation etc.
For example, capitals would be mapped to ‘X’, lower case to ‘x’, digits to ‘d’, and punctuation to itself. So “CPA1” would be mapped to “XXXd”, and “mRNA” to “xXXX”.
Strings were also shortened, the first two letters and last two letters were encoded by the scheme, but longer words would be truncated. For all the characters in the middle you’d just include the sets of the different character classes that were included.
So you’d end up with dense representations that extract useful lexical features of the word.
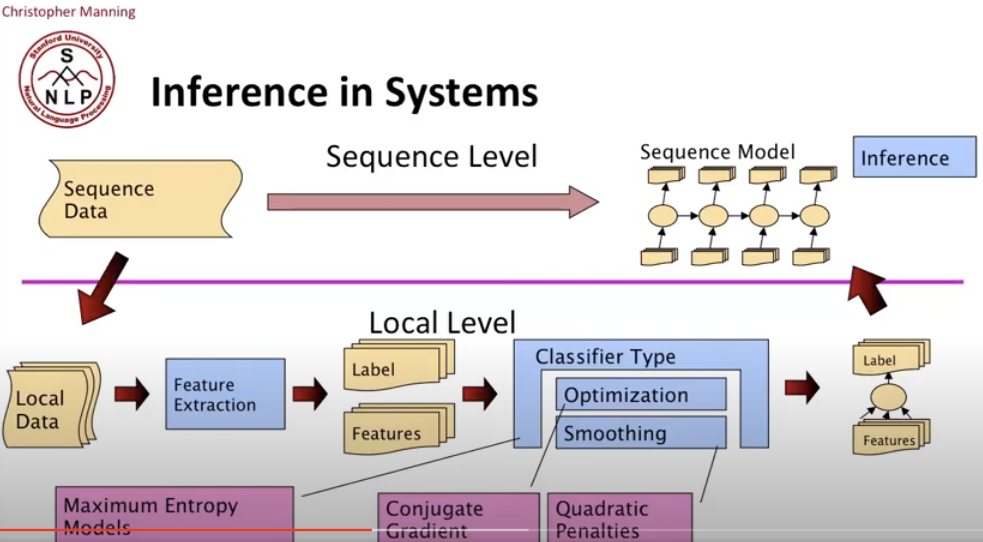
Maximum Entropy Sequence Models
Many problems in NLP have data which is a sequence of something. Could be characters, words, phrases, paragraphs, lines, sentences. We can think of our task often as labeling each item in the sequence.
Sometimes even problems that don’t initially look like sequence labeling can be modeled like that. For example we have a FAQ page of a website and we want to parse its structure so we understand which sections are questions and which answers. We can treat this as a sequence labeling task where we assing the labels ‘Q’ or ‘A’ to paragraphs.
We can tackle this through putting maximum entropy models into a sequence model.
For a Conditional Markov Model (CMM) aka a Maximum Entropy Markov Model (MEMM), the classifier makes a single decision at a time, conditioned on evidence from observations and previous decisions.
A larger space of sequences is usually explored via search. For example consider the following local context, where we want to assign a POS label to token 0:
| -3 | -2 | -1 | 0 | +1 |
|---|---|---|---|---|
| DT | NNP | VBD | ??? | ??? |
| The | Dow | fell | 22.6 | % |
To do this we’ll be able to use as features of the surrounding words. But we’ll also be able to use as features the labels assigned for previous tokens, say 2 back.
In a greedy approach, once a label is assigned it’s not modifiable, the classifier just keeps going along the sequence.
More commonly though, you want to be able to revisit the classification of a prior element in the sequence as a result of a subsequent decision. There are different methods to do that.
One is beam inference. Here, at each position rather than just keep one label we keep the top k complete sequences. We then extend each of these sequences in a local way, the extensions compete for the top k slots in the next position.
The advantages of beam inference are that it’s fast. Beam size of 3-5 are almost as good as exact inference in many cases. And it’s easy to implement, we don’t need dynamic programming.
The disadvantage is that it is inexact, the globally best sequence can fall of the beam.
Another approach, that is guaranteed to return the global optimum, is Viterbi Inference, which proceeds through dynamic programming or memoization. It requires a small window of state influence (eg past two states are relevant).
The main advantage is its exactness, the disadvantage is that it is harder to implement long-distance state-state interactions (but then beam search doesn’t often allow long-distance sequence resurrection anyway).
Conditional Random Fields
An alternative sequence model to MEMMs is Conditional Random Fields or CRFs. This is a whole-sequence conditional model, rather than a chaining of local models.
It can be expressed as the equation of:
\[P(c | d, \lambda) = \frac{\exp \sum_i \lambda_i f_i(c,d)}{\sum_{c’}\exp\sum_i \lambda_i f_i(c’,d)}\]
This looks like it would be impossible to deal with, given the space of whole sequence inputs is potentially infinite, and the space of whole sequence class outputs is exponential.
But if the features \(f_i\) remain local, the conditional sequence likelihood can be calculated exactly using dynamic programming.
Training is slower, but CRFs avoid causal-competition biases, so have theoretical advantages over MEMMs.
At the time of the lectures (2012) these were seen as state of the art, but performed much the same as MEMMs in practice.